
 B.Eng., Zhejiang University (2027)
B.Eng., Zhejiang University (2027)Hi! I'm Zijie, a junior CS student at Zhejiang University in Turing Class.
My research interst lies in the intersection of deep generative models and AI4Sciences,
aiming at interpreting the biological world.
Previously I focus on molecular design and optimization with specific constraints.
I hope to leverage AI to pioneer innovative therapeutics that change the world.
I was fortunate to start my research journey at CADD Lab under the supervision of
Prof. Hou Tingjun. I also feel lucky to be advised by Prof. Jiang Kaiyi from Princeton University.
I'm also a member of Zhejiang University Orchestra playing 🎷, was PG for CKC college🏀 and a fan of Leo Messi and Stephen Curry.
Warning
Problem: The current name of your GitHub Pages repository ("Solution: Please consider renaming the repository to "
http://".
However, if the current repository name is intended, you can ignore this message by removing "{% include widgets/debug_repo_name.html %}" in index.html.
Action required
Problem: The current root path of this site is "baseurl ("_config.yml.
Solution: Please set the
baseurl in _config.yml to "Education
-
 Zhejiang UniversityB.Eng. in Computer Science(Honor)Sep. 2023 - Jul. 2027
Zhejiang UniversityB.Eng. in Computer Science(Honor)Sep. 2023 - Jul. 2027
Experience
-
 Princeton University Jiang LabResearch InternSep. 2025 - present
Princeton University Jiang LabResearch InternSep. 2025 - present -
 Zhejiang University CADD LabResearch InternJun. 2024 - present
Zhejiang University CADD LabResearch InternJun. 2024 - present
Honors & Awards
-
Zhejiang University First-Class Scholarship2024 2025
-
Zhejiang University Excellent Student2024 2025
News
Selected Publications (view all )
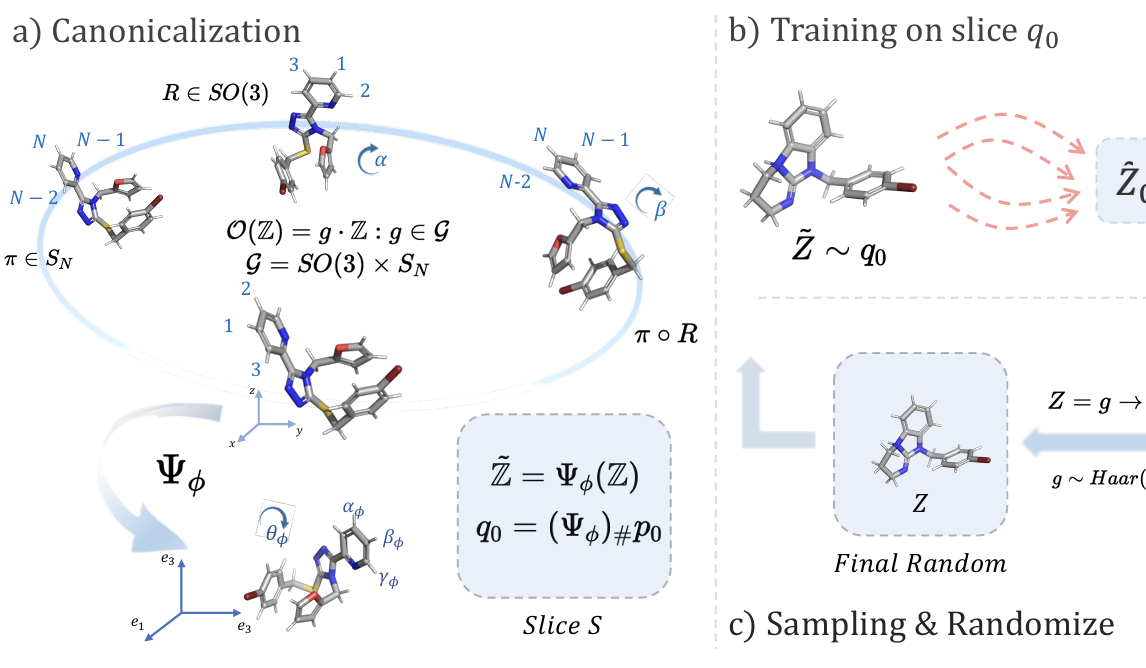
Rethinking Diffusion Models with Symmetries through Canonicalization with Applications to Molecular Graph Generation
Cai Zhou*, Zijie Chen*, Zian Li, Jike Wang, Kaiyi Jiang, Pan Li, Rose Yu, Muhan Zhang, Stephen Bates, Tommi Jaakkola (* equal contribution)
Preprint 2026
Canonicalization framework for symmetry-invariant generative modeling, that removes symmetry-induced ambiguity in diffusion training and achieving state-of-the-art performance in 3D molecule generation.
Rethinking Diffusion Models with Symmetries through Canonicalization with Applications to Molecular Graph Generation
Cai Zhou*, Zijie Chen*, Zian Li, Jike Wang, Kaiyi Jiang, Pan Li, Rose Yu, Muhan Zhang, Stephen Bates, Tommi Jaakkola (* equal contribution)
Preprint 2026
Canonicalization framework for symmetry-invariant generative modeling, that removes symmetry-induced ambiguity in diffusion training and achieving state-of-the-art performance in 3D molecule generation.
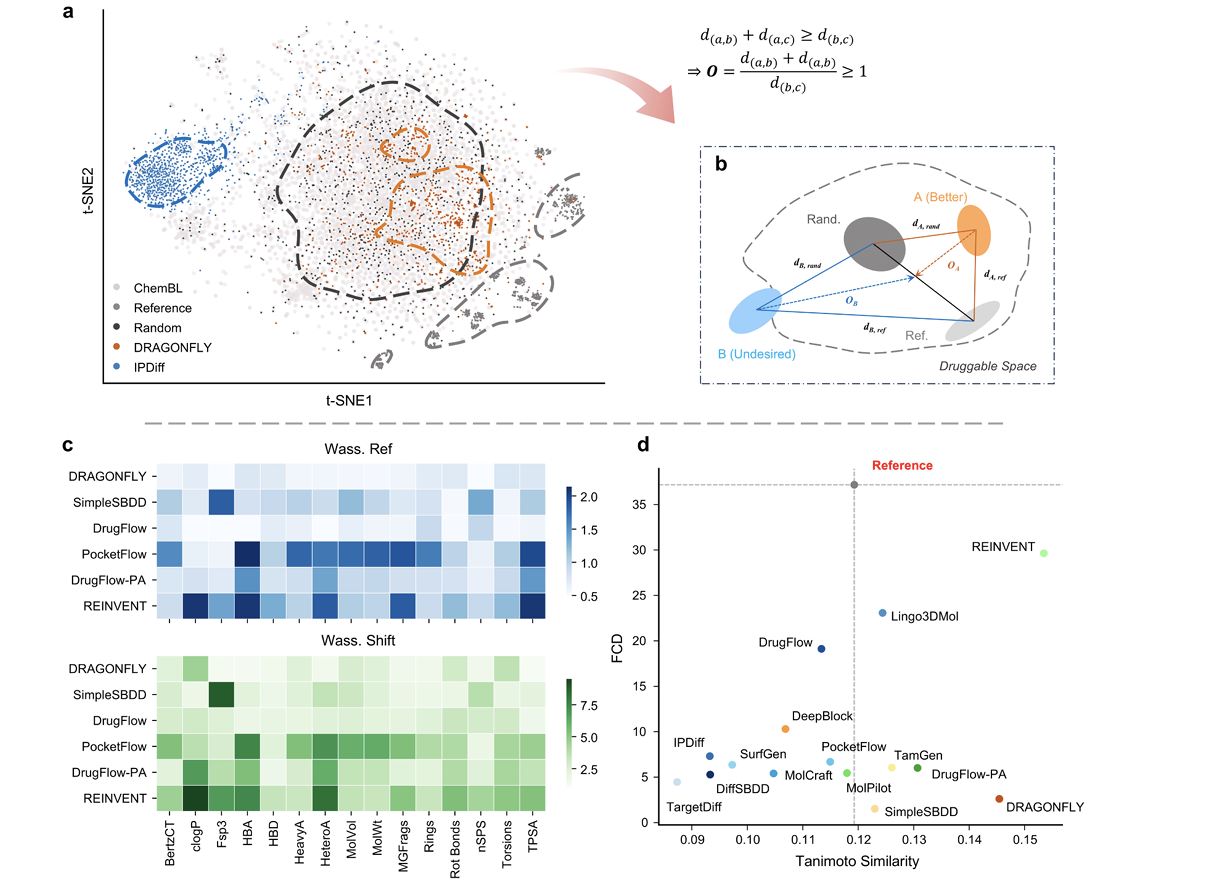
Revisiting Target-Aware de novo Molecular Generation with TarPass: Between Rational Design and Texas Sharpshooter
Rui Qin*, Zijie Chen*, Yurong Li, Meijing Fang, Longji Shen, Yilong Su, Odin Zhang, Qinghan Wang, Qun Su, Jike Wang$^{\dagger}$, Tingjun Hou$^{\dagger}$, Yu Kang$^{\dagger}$ (* equal contribution)
Preprint 2026
TarPass, a rigorous benchmark for target-aware de novo molecular generation, building a scalable evaluation pipeline and uncovering a critical gap between favorable docking metrics and faithful protein–ligand interaction modeling.
Revisiting Target-Aware de novo Molecular Generation with TarPass: Between Rational Design and Texas Sharpshooter
Rui Qin*, Zijie Chen*, Yurong Li, Meijing Fang, Longji Shen, Yilong Su, Odin Zhang, Qinghan Wang, Qun Su, Jike Wang$^{\dagger}$, Tingjun Hou$^{\dagger}$, Yu Kang$^{\dagger}$ (* equal contribution)
Preprint 2026
TarPass, a rigorous benchmark for target-aware de novo molecular generation, building a scalable evaluation pipeline and uncovering a critical gap between favorable docking metrics and faithful protein–ligand interaction modeling.